Plant pathogens trigger changes in host plants that allow them to cause disease. Understanding which molecules pathogens use to do this (termed effectors), how they allow infection to take place, and how they are detected by plants has important implications for our understanding of plant disease.
SEFARI research in this area has allowed major improvements in our understanding of how the plant immune system works, how the immune system and other plant processes are linked, and how resistance that targets essential effectors can be identified. Our fundamental work in this area has led to collaborations with industrial partners, underpinning the development of new control strategies that target key infection processes.
Stage
Directory of Expertise
Purpose
It is estimated about 40% of global crop production is lost to pests and diseases. Within Scotland, a range of pathogens including fungi, nematodes, oomycetes (which cause potato late blight and raspberry root rot) and aphids damage key crops such as barley, potato and soft fruits. To develop new control strategies based on host resistance, integrated pest management or new agrochemicals, we need to understand how pathogens infect plants and how this process can be disrupted.
Although the pathogens that we aim to control are incredibly diverse, all need to manipulate host plant biochemistry for successful infection, and all do this using secreted molecules called effectors. Effectors may switch off the host immune system or cause changes in the plant so that it provides the pathogen with the resources it needs for development. SEFARI scientists have identified effectors from different crop pathogens and are working to understand the plant processes that they target. This knowledge informs novel and specifically tailored pathogen control strategies for sustainable crop production.
Results
We have used genome sequencing and analysis to allow us to identify effectors in our plant pathogen research. Sequencing has become much cheaper and faster in recent years, allowing us to sequence DNA of different strains of our pathogens or to specifically target sequencing on effectors. By comparing the sequences of different pathogens we can identify effectors that are present in each strain and- that are therefore likely to be essential for infection of the host plant. By ‘knocking out’ individual effectors, using gene silencing approaches to reduce expression of the relevant genes, we have also been able to identify those that play a critical role in pathogen biology. Through targeting, and focusing on, these key molecules, we can develop and understand the most durable forms of host resistance.
Using different approaches, we can identify the plant proteins that are the targets of the pathogen effectors. Our analysis has shown us that pathogens target incredibly diverse processes in plants in order to stop their defences from operating properly. Effectors from all the pathogens we have studied target important signalling pathways to prevent them from functioning normally. For example, the processes of plant defence and growth are mutually exclusive – only one can operate at a time and each suppresses the other. Oomycete (fungus-like) pathogens including the late blight pathogen Phytophthora infestans, take advantage of this by activating plant growth and therefore inhibiting defence responses. Meanwhile, nematodes (small soil dwelling parasitic roundworms) produce effectors that target host biochemistry so that the plant produces high levels of specific nutrients that are required by the nematode.
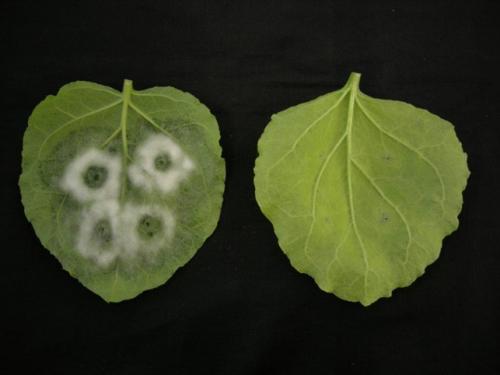
Figure: A microscope image of the late blight pathogen infecting a plant leaf. Plant cell components are highlighted in blue and yellow, and the body of the pathogen is in green. An effector is highlighted in magenta and from this we can see it is specifically produced at finger-like projections from the pathogen body which penetrate the plant cells to deliver effectors. The white bar indicates ten micrometres in length.
Benefits
This research has generated significant additional external funding, adding value to our Scottish Government funded research, and has led to high quality publications (listed below).
Funding from industry, for example, has enabled work on late blight which has led to the identification of a completely new mechanism that the pathogen uses to secrete effectors into the plant. The possibility of targeting this process for development of new agrochemicals is currently being investigated in a UK Research Innovation (UKRI) funded project partially funded by a large agro chemical company.
By understanding which processes in the plant are targeted by the pathogens, we have a better understanding of how the plant immune system functions and how the immune system and other essential plant processes are interlinked. This allows us to understand the wider consequences of approaches aimed at making plants more resistant to pathogens. For example, we can now better predict how making changes to these pathways, either through breeding new varieties or the application of agrochemicals, may tip the delicate balance between plant yield and loss due to pests and diseases.
Effectors are part of the pathogen that is recognized when a resistant host response is triggered. A recognized effector renders a pathogen unable to infect their host plant and is called an avirulence gene. We have collaborations with breeding companies that aim to identify durable resistance genes. We now systematically screen wild species of cultivated crops for the presence of disease resistance genes that are attuned to detecting critical pathogen effectors and induce a resistance response in the plant. Furthermore, we can direct resistance breeding efforts to only combine complimentary resistances, e.g. those genes that target multiple diverse effectors to ensure more durable pathogen control. Large, international potato breeders and processors are joining forces with SEFARI scientists to capitalise on these advancements.
Project Partners
Staff involved in this project have joint appointments with the University of Dundee Division of Plant Sciences and the University of St Andrews School of Biology