Beef production is very important for Scotland's economy (economic output of £851 million in 2017) and for providing the high quality and iconic, ‘Scotch Beef’ brand. Feed for livestock represents 70-80% of the variable costs of beef production and an improvement of feed conversion efficiency is needed to keep the beef industry economically viable. Furthermore, a reduction in carbon footprint has to be achieved in order to minimize environmental impact. This prize-winning research showed that both improvement in feed conversion efficiency and mitigation of methane emissions is achievable through breeding, by using information from microbes and their genes present in the rumen (the first of cow’s four ‘stomachs’). This breeding strategy is cost-effective because it avoids the need to make expensive measurements of feed conversion efficiency and methane emissions.
Stage
Directory of Expertise
Purpose
By 2050, the Food and Agriculture Organisation of the United Nations has reported that the human population will increase to over 9 billion people, and in the same time-frame, global meat consumption is projected to increase by 76%.
Improved feed conversion efficiency, that means producing more meat per kg of feed consumed, will be an important part of the response to this challenge. However, intensive food production puts a strain on the environment, while there is also a need to produce more food in an ethical manner.
The rumen microbial ecosystem comprises a very dense community of different bacteria, Archaea, Protozoa and Fungi. This ecosystem within the rumen is particularly important in cattle due to its ability to convert indigestible fibrous plant material (e.g. grass) into absorbable nutrients used to produce high quality food, such as meat and milk, for human consumption.
As a by-product of the microbial conversion of feed (in particular fibrous grass), the rumen microbial Archaea population produces methane, which is expelled through mouth and nose into the environment. Methane is a greenhouse gas (GHG) with global warming potential 28-times that of carbon dioxide and cattle are a major source of anthropogenic methane emissions.
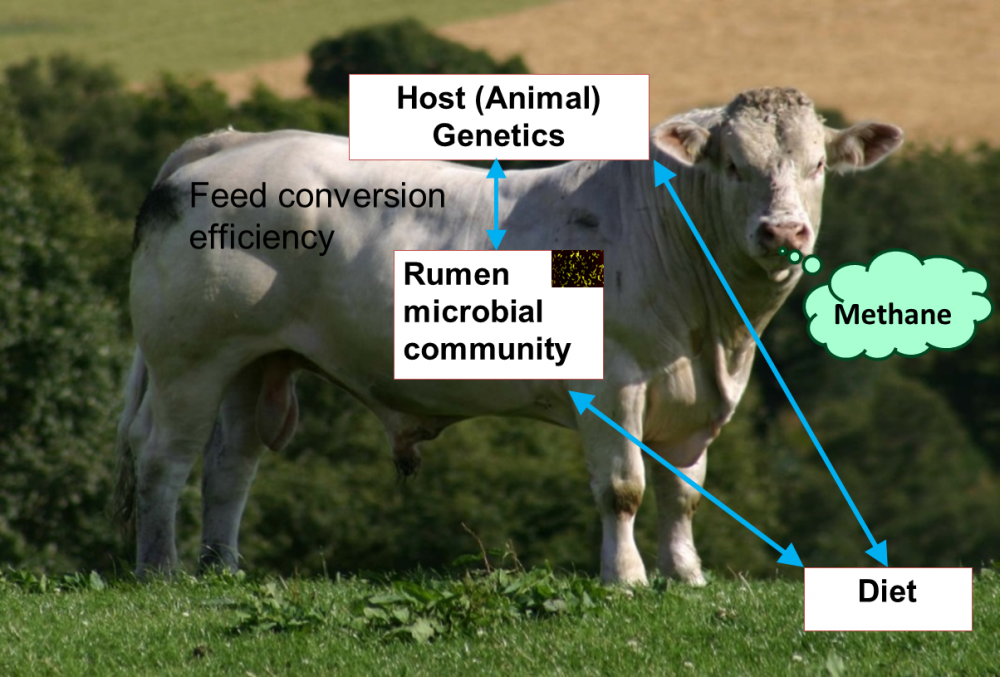
In order to address food security, as well as economic and environmental impacts of food production, sustainable intensification has been suggested. For farm animals, the genetic improvement of feed conversion efficiency is deemed the most important way to achieve it.
Finding means to mitigate methane emissions from cattle and to improve their feed conversion efficiency are the main emphasis of the Scottish Government-funded research in Theme 2 of ‘Productive and Sustainable Land Management and Rural Economics’ in the current Strategic Research Programme, 2016-2021.
Results
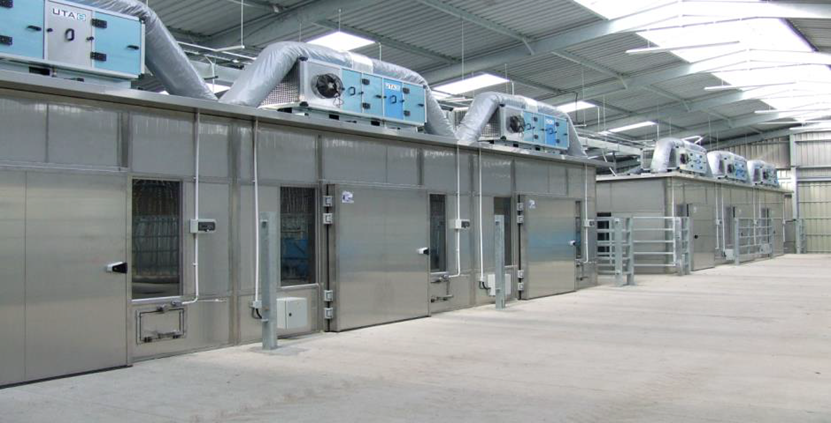
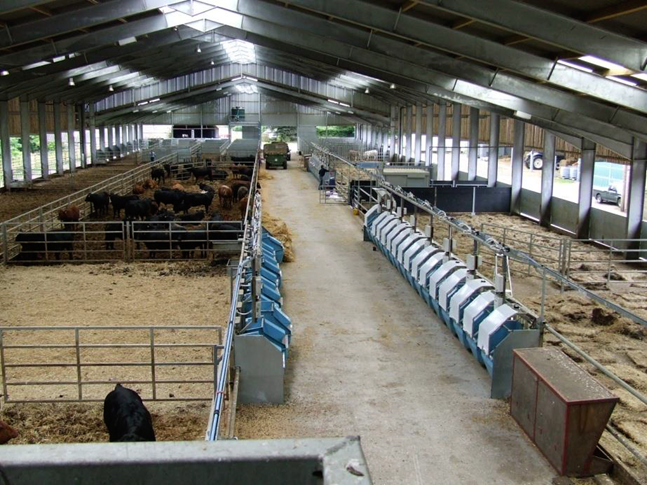
At SRUC’s Beef and Sheep Research Centre, we recorded the feed intake of beef cattle using electronic feeders and subsequent methane emissions using respiration chambers.
In addition, we obtained samples of rumen contents, either using a stomach tube on live animals or directly from the rumen of slaughtered animals in the abattoir. We sequenced microbial DNA extracted from the rumen samples to identify the relative abundance of different microbes (e.g. bacteria and Archaea) and microbial genes.

We found that the relative abundance of Archaea, which are the main organisms producing methane in the rumen of cattle, are a good predictor of methane emissions. However, the relative abundance of key microbial genes provided an even better predictor for methane emissions.
Most of the identified microbial genes are involved in the metabolism of methane or pathways providing substrates (e.g. hydrogen) for methane metabolism. We also revealed that the relative abundance of methanogenic Archaea is controlled by the host animal’s genetics. This means that breeding can be used to reduce the methanogenic Archaea and thus methane emissions.

The relative abundance of different microbial genes was also found to be a good predictor of feed conversion efficiency so that both methane emissions and feed conversion efficiency can be predicted from different microbial genes in the same sample. The relationship between the abundance of microbial genes and methane emissions was independent of the diet so that breeding could be combined with dietary intervention to cumulatively mitigate methane emissions
Benefits
This research provides new breeding strategies to improve the sustainability of Scottish beef production in terms of efficiency and mitigation of GHG emissions.
During a 12-week feeding period, we calculated feed cost savings of £23/head of cattle between the most feed efficient third and least efficient third in a Stabiliser cattle population. For 388,700 prime cattle slaughtered in Scottish abattoirs during 2017, this would mean a cost reduction of £8.9 million, considering only 3 months of the growing-finishing period.
The use of microbiome information to predict feed conversion efficiency will save the large cost of recording feed intake and growth in test stations. It is, therefore, a cost-effective and practical strategy for genetic improvement of feed conversion efficiency.
Since beef cattle have never been selected for reduction in methane emissions before, we obtained a large variation in methane emissions from 170g/day to 330g/day among beef cattle of the same breed offered the same forage-based diet.
This large variation of methane emissions, and the host genetic control of methane-producing Archaea, will result in a selection response for reduction of methane emissions (expected to be about 3% of the mean per year).
Measuring methane emissions in respiration chambers for breeding purposes is very accurate but too costly and cannot provide the number of animals neccessary to be recorded for accurate breeding.
In contrast, a more cost-effective method to predict methane emissions is by using the relative abundance of different microbial genes, based on rumen samples taken using a stomach tube on live animals or directly from the rumen of slaughtered animals. This method will provide a dataset of predicted methane emissions from a large number of individual animals to be used to make accurate breeding decisions to select animals emitting less methane.
More broadly, our research suggests that rumen microbial genes are excellent biomarkers for complex traits linked to other important issues - such as animal health and welfare, meat fatty acid profiles (healthy diets for people) and antimicrobial resistance.